GPU
GPU stands for "Graphic Processing Unit". If you have access to a dedicated GPU device, the iteration procedure will run much faster. While Dynamo can run its GPU functionalities in most Nvidia cards, a real advantage will be measured for those cards that are conceived for scientific computation. The Tesla series and the Titan series are examples of this.
Contents
Installation
A regular distribution of Dynamo will include GPU precompiled GPU executables. As they need to be linked to libraries that might not be present in your system, it is advised that you recompile them:
After untarring the tar package in location DYNAMO_ROOT, go to
cd DYNAMO_ROOT/cuda
and make certain that you have CUDA active in your shell, for instance look for the NVIDIA nvcc compiler:
which nvcc
If this is succcessful run
source config.sh
This will automatically edit the makefile file in the folder, informing it on the location of CUDA in your system. In order to be sure, you can also edit the makefile text file and make sure that the line
CUDA_ROOT=/usr/local/cuda-7.5
have been edited to the correct CUDA libraries on your system. The CUDA_ROOT variable expected here can be infered from the path to the detected nvcc compiler by <CUDA_ROOT>/bin/nvcc
Once your makefile is correctly formated, you can just type:
make clean
to delete the executables already shipped in the distribution (which might not be compatible with your libraries), and then
make all
to recompile the executables in your system.
CUDA libraries
You need to have CUDA installed on your system. This might require coordinating with your system administrator.
Versions
Dynamo has been tested with most CUDA versions. CUDA 7.0 was found to show problems in the Fourier transform libraries: don't use it. We advice to use the highest available CUDA libraries, at least CUDA 7.5 per 18/04/2016.
Testing your system
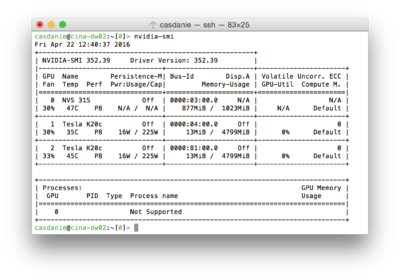
In the Linux shell type: nvidia-smi to get a list of the GPU devices that your system is seeing, their status and the jobs currently running. This order also shows the device number assigned by the system to each device, which you need to enter in a Dynamo project through the parameter gpu_identifier_set
Speed up
This iis of course highly dependent on the system: which GPUs you are comparing against which CPU cores.
It also depends on the particle size and on the number of angles scanned per particle (higher number will yield higher speed up factors, i.e., the more intensive a computation, the more favorable for GPU)
Using the GPU in an alignment project
Your execution environment should include a LD_LIBRARY_PATH environment variable that includes the location of the CUDA libraries. You probably need to inform your UNIX shell with a command similar to this:
export LD_LIBRARY_PATH=$LD_LIBRARY_PATH:/usr/local/cuda75/lib
Here, replace with the location of the CUDA libraries in your system. If you have several CUDA installations, choose the one that you used to compile Dynamo in your system. Also if you are going to work under Matlab you should update your LD_LIBRARY_PATH variable in the Linux shell before starting Matlab.
With the dcp GUI
Go to the Computing environment GUI. Select 'matlab gpu' or 'system gpu' Set the gpu identifier set field to the device numbers given to your GPUs by the operative system. You can select a subset of the available GPUs. If you have a GPU that controls the screen display, you can technically use it for GPU computations, but it is not advisable, as it will be typically much slower than GPUs intended for computation.
With the command line
There are two project parameters related the the GPU: the destination parameter and the gpu_identifier_set parameter. The destination must be set to matlab_gpu (to run projects inside Matlab) or to system_gpu
Checking your system
- Main article: GPU identifiers
In a linux machine nvidia-smi will give you a status summary of the installed, reachable GPUs. In the example image, you would use an identifier_set of 1,2 to. Alternatively, you could run different projects on device 1 and 2. In any case, device 0 is not intended for GPU computing.
GPUs in classification
GPUs in classification are only available in the context of MRA alignment. Multireference alignment inherits the "GPU-friendliness" of single reference alignment: the cc of many rotations of the template against the particle can be computed inside the GPU without any transfer of data into the CPU or the hard disk.
In PCA, the situation is totally different: computation of the cross correlation matrix is not a compute-intensive process, but rather a data-intensive process: we don't rotate several times the same data particle. Particles need to be read from disk, rotated and compared against other particles (which have undergone the same process). For this reason, a GPU version of cc-matrix computation for PCA analysis is not available, as it would not provide any speed-up in comparison with a CPU application. The only effective way to speed up PCA is the use of parallel CPU computing.