Uncategorized files
Jump to navigation
Jump to search
Showing below up to 500 results in range #1 to #500.
View (previous 500 | next 500) (20 | 50 | 100 | 250 | 500)
- 3d-silce-along-with-active-pool-model-dmmpp.png 1,709 × 1,584; 481 KB
- 3d-silce-along-with-active-pool-model.png 1,772 × 1,638; 487 KB
- ABPcommandCoarse.png 1,872 × 1,125; 464 KB
- ABPdata.png 1,250 × 1,125; 468 KB
- ABPfineComparison.png 2,350 × 1,125; 286 KB
- ABPfineResult.png 2,073 × 1,125; 447 KB
- ABPmanualABP.png 1,405 × 1,125; 262 KB
- ABPmanualABPTemplate.png 1,870 × 1,226; 311 KB
- ABPmanualABPinput.png 1,944 × 1,002; 303 KB
- ABPmanualCheck.png 1,169 × 1,125; 172 KB
- ABPmanualCreate.png 2,142 × 1,125; 289 KB
- ABPmanualEnvironment.png 2,071 × 1,125; 314 KB
- ABPmanualFSC.png 1,250 × 1,125; 256 KB
- ABPmanualFSCAttained.png 1,404 × 1,125; 236 KB
- ABPmanualFSCInteractive.png 1,249 × 1,125; 294 KB
- ABPmanualFSCInteractiveIte1.png 2,245 × 1,125; 736 KB
- ABPmanualFSCInteractiveLastIteration.png 2,191 × 1,125; 726 KB
- ABPmanualFSCinitial.png 2,777 × 955; 365 KB
- ABPmanualMasks.png 2,000 × 1,175; 311 KB
- ABPmanualNumericalParameters.png 1,667 × 1,344; 297 KB
- ABPmanualParameters.png 2,177 × 1,125; 362 KB
- ABPmanualParametersEdit.png 2,406 × 886; 183 KB
- ABPmanualParticles.png 1,933 × 1,125; 269 KB
- ABPmanualTable.png 2,097 × 1,071; 301 KB
- ABPmanualTemplate.png 2,144 × 1,254; 298 KB
- ABPmanualUnfold.png 1,169 × 1,125; 171 KB
- AWF GB definitions.png 865 × 480; 79 KB
- Activation standalone.png 1,074 × 659; 64 KB
- Adapted-mask-sta.png 2,983 × 1,485; 562 KB
- Adapted-mask.png 2,983 × 1,485; 557 KB
- AddStorageMain.png 1,500 × 893; 332 KB
- Ali mask Z.png 821 × 837; 31 KB
- Ali mask y.png 821 × 837; 24 KB
- Ali ts1.png 1,620 × 1,414; 2.81 MB
- AlignCommandLine autocompletionOfFurtherFieldsOfUserObject.png 570 × 380; 91 KB
- AlignCommandLine autocompletionOfUserObject.png 4,190 × 1,993; 1.09 MB
- AlignCommandLine autocompletionOfUserObjectEMBO2021.png 4,190 × 1,332; 922 KB
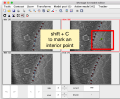
- AlignCommandLine clickAroundTenMarkers.png 1,344 × 1,362; 660 KB
- AlignCommandLine createANewModel.png 1,532 × 354; 184 KB
- AlignCommandLine dslicesOnBinnedGoldBeads.png 1,344 × 1,210; 405 KB
- AlignCommandLine dslicesOnFullSizedGoldBeads.png 1,344 × 1,210; 640 KB
- AlignCommandLine dtmsliceAfterContrastCorrection.png 1,428 × 1,254; 437 KB
- AlignCommandLine dtmsliceContrastControl.png 226 × 250; 14 KB
- AlignCommandLine dtmsliceOfBinnedReconstruction.png 2,772 × 1,672; 748 KB
- AlignCommandLine exportATableIntoTheMatlabWorkspace.png 1,424 × 894; 491 KB
- AlignCommandLine fullProjectionOfBinnedReconstruction.png 1,344 × 1,362; 654 KB
- AlignCommandLine mainClickToFixThe3dCoordinates.png 1,344 × 1,362; 444 KB
- AlignCommandLine rightClickOnAPoint.png 1,344 × 1,362; 656 KB
- AlignCommandLine showAFullProjection.png 938 × 302; 67 KB
- AlignCommandLine viewOfTiltLines.png 1,344 × 1,210; 460 KB
- AlignCommandLine xViewOfClickedPoint.png 1,344 × 1,362; 443 KB
- AlignGUI DetectionParameters.png 1,136 × 1,148; 181 KB
- AlignGUI EachPatchWillBeAPutativeGoldBead.png 1,135 × 197; 71 KB
- AlignGUI ExtractedRefinedPeaks.png 1,128 × 1,138; 233 KB
- AlignGUI ExtractingPeaks.png 1,132 × 1,140; 243 KB
- AlignGUI YouMightNeedToActivateTheBandpassToSeeTheVLP.png 1,846 × 1,670; 690 KB
- AlignGUI alignmentGUIBeforeCompletionOfAnyStep.png 1,344 × 1,170; 353 KB
- AlignGUI allStepsOfRefinementAreaAreBeingComputed.png 1,129 × 903; 81 KB
- AlignGUI areaForCreationOfAnAlignedTiltSeries.png 1,344 × 1,170; 350 KB
- AlignGUI betterFiitingIsAttained.png 920 × 872; 269 KB
- AlignGUI binnedAlignedTiltStack.png 1,846 × 1,672; 1.04 MB
- AlignGUI binnedWeightedBackProjectionReconstruction.png 1,478 × 1,358; 617 KB
- AlignGUI checkMarkerPositionsAfterClustering.png 1,375 × 122; 66 KB
- AlignGUI checkObservationCloudAfterSelectionOrClustering.png 1,350 × 164; 122 KB
- AlignGUI chooseAVisualization.png 2,106 × 1,146; 407 KB
- AlignGUI cloudOfUnindexedCcPeaks.png 1,344 × 1,210; 500 KB
- AlignGUI coloringOptionForScatteredPlots.png 1,336 × 826; 186 KB
- AlignGUI completedStepsBecomeBoldFontSize.png 1,344 × 1,170; 354 KB
- AlignGUI computingRawCCMatrixt.png 1,134 × 1,138; 246 KB
- AlignGUI creatingAWorkflow.png 1,458 × 440; 142 KB
- AlignGUI creationOfWorkflowGUITakesSomeSeconds.png 628 × 378; 87 KB
- AlignGUI differentPlottingOptions.png 1,424 × 1,634; 470 KB
- AlignGUI eachPointRepresentsACcPeak.png 1,344 × 1,210; 457 KB
- AlignGUI exampleOfFailedSearch.png 1,718 × 1,422; 562 KB
- AlignGUI featurePlotterAllowsCheckingOfTheFeatureValuesOnTheWholeSet.png 1,104 × 158; 49 KB
- AlignGUI inputWorkflowParameters.png 1,208 × 778; 200 KB
- AlignGUI markerPositionsAfterClustering.png 1,718 × 1,330; 853 KB
- AlignGUI markerSetsCanBeViewedAsCloudOfPoints.png 1,118 × 210; 77 KB
- AlignGUI noteWorkingMarkersCanBeRestoredToTheResultOfAPreviousStep.png 1,285 × 316; 193 KB
- AlignGUI observationCloudHasBeenThinned.png 1,344 × 1,210; 522 KB
- AlignGUI occupancyAfterTiltGapFiller.png 1,344 × 1,210; 386 KB
- AlignGUI occupancySchemeOfTheResultsOfAStep.png 1,377 × 266; 111 KB
- AlignGUI openTomoshowOnTheFullSizedMatrixToCheckTheSizeOfAGoldBead.png 1,846 × 1,656; 1.85 MB
- AlignGUI parametersForIterativeReindexing.png 1,344 × 1,470; 463 KB
- AlignGUI parametersForTiltGapFiller.png 1,344 × 1,710; 533 KB
- AlignGUI parametersOfTiltExtensor.png 1,344 × 1,650; 516 KB
- AlignGUI passingTheCursorOnTheGUIElementsShowsAHelpElement.png 1,035 × 679; 204 KB
- AlignGUI patchGalleryForFeatures.png 1,139 × 314; 162 KB
- AlignGUI reconstructionArea.png 1,198 × 1,182; 247 KB
- AlignGUI refinedCCMatrix.png 1,130 × 1,138; 237 KB
- AlignGUI residualsAtEachObservation.png 789 × 737; 113 KB
- AlignGUI resultOfIterativeReindexing.png 1,344 × 1,210; 384 KB
- AlignGUI resultOfTrailExtensor.png 1,718 × 1,330; 845 KB
- AlignGUI resultsOfTestingOnACentralImage.png 1,344 × 1,262; 1.04 MB
- AlignGUI resultsOfTiltGapFiller.png 1,718 × 1,330; 845 KB
- AlignGUI roughAlignmentSelectsManyPotentialTrails.png 1,344 × 1,210; 384 KB
- AlignGUI scatter2dOption.png 1,424 × 1,034; 319 KB
- AlignGUI showsTheDetectedSpotsOnTheVideoGUI.png 1,164 × 200; 75 KB
- AlignGUI someOfTheNewTrailsHaveEmptySpaces.png 1,344 × 1,210; 395 KB
- AlignGUI staticVideoAnnotationsCannotBeEdited.png 1,718 × 1,330; 1.48 MB
- AlignGUI tabForRefinement.png 1,344 × 1,170; 365 KB
- AlignGUI theFullSizeReconstructionIncludesOnlyTheObjectOfInterest.png 1,508 × 1,350; 699 KB
- AlignGUI theGalleryAllowsRelatingAppearanceOfGoldBeadAnValueOfTheFeatures.png 1,718 × 1,422; 584 KB
- AlignGUI thePinToolIsUsefulForLocatingIsolatedPoints.png 1,488 × 1,284; 281 KB
- AlignGUI thePlusIconAtCornerShowsTheValuesOfTheParametersOfAGivenStep.png 1,344 × 1,350; 416 KB
- AlignGUI theRulerToolAllowsToMeasureInPixelsTheDiameterOfAGoldBead.png 1,846 × 1,656; 646 KB
- AlignGUI theSetOfCcPeaksIsShownAsAMontage.png 1,718 × 1,422; 563 KB
- AlignGUI toolboxOptionForTestingTheParametersOnACentralImage.png 1,494 × 1,174; 355 KB
- AlignGUI trailExtensorAddsAFewNewTrails.png 952 × 216; 122 KB
- AlignGUI trailsWithFewMarkersAreThenRejected.png 1,344 × 1,210; 385 KB
- AlignGUI trimmingOfMarkersCanBeAccessedThroughTheScissorIcon.png 795 × 543; 155 KB
- AlignGUI visualizationOfAlignedTiltStack.png 1,686 × 228; 86 KB
- AlignGUI weCheckTheDistanceOfTheObjectOfInterest.png 1,488 × 1,284; 283 KB
- AlignGUI weHave66TracesAfterGapFiller.png 968 × 945; 308 KB
- AlignGUI wePutAPinInTheCenterOfAnObjectOfInterest.png 1,846 × 1,670; 1.02 MB
- Align gui check marker clustered position.png 2,118 × 297; 158 KB
- Align gui check observation cloud after selection or clustering.png 2,104 × 257; 128 KB
- Align gui cloud unindexed cc peaks.png 1,137 × 982; 175 KB
- Align gui detected spots on video gui.png 2,024 × 365; 145 KB
- Align gui detection parameters.png 1,344 × 1,362; 411 KB
- Align gui marker sets as point cloud.png 2,033 × 371; 149 KB
- Align gui observation cloud.png 1,114 × 990; 152 KB
- Align gui occupancy scheme.png 2,032 × 358; 156 KB
- Align gui parameters tilt extensor.png 1,211 × 1,504; 226 KB
- Align gui patch as putative gold bead.png 2,028 × 362; 143 KB
- Align gui patch gallery for features.png 1,872 × 408; 180 KB
- Align gui point representing cc peak.png 1,097 × 849; 156 KB
- Align gui traces after gap filler.png 1,438 × 1,527; 427 KB
- Align gui working markers restored.png 1,905 × 422; 182 KB
- Alignment-project-main-block-iterative-workflow-sta.png 1,870 × 911; 440 KB
- Alignment-project-main-block-iterative-workflow.png 1,948 × 1,015; 447 KB
- An example of a gain reference.png 1,458 × 822; 507 KB
- AngularGridAnglePlot.png 1,120 × 840; 51 KB
- AngularGridFlip.png 1,120 × 840; 48 KB
- Astigmatism conventions.jpg 563 × 581; 74 KB
- Av ali ts1.png 1,616 × 1,412; 2.99 MB
- Average-bin-2-from-seed-oversampling-dmmpp.png 1,166 × 1,163; 166 KB
- Average-bin-2-from-seed-oversampling.png 1,185 × 1,184; 46 KB
- Average-overplot-hexagonal-lattice-measurements.png 904 × 850; 272 KB
- Average hiv.jpg 1,275 × 424; 289 KB
- BasicLaceyStepBinarization.png 294 × 288; 5 KB
- BasicLaceyStepDilation.png 291 × 290; 33 KB
- BasicLaceyStepIntensitySelection.png 293 × 288; 84 KB
- BasicLaceyStepOriginal.png 293 × 289; 84 KB
- BasicLaceyStepReordering.png 292 × 292; 90 KB
- BasicLaceyStepSobel.png 297 × 293; 128 KB
- BasicLaceySupportGUIFullView.png 600 × 1,000; 540 KB
- BasicLaceySupportGUIGlobal.png 608 × 1,001; 492 KB
- BasicLaceySupportingGUILocal.png 608 × 1,001; 591 KB
- BasicLaceyWorkflowGUI.png 600 × 970; 48 KB
- Borrelia set average.png 667 × 628; 155 KB
- Borrelia set particle example.png 887 × 452; 111 KB
- CTF problems in cryoem.png 1,122 × 894; 1.34 MB
- Catalogue manager.png 1,350 × 743; 140 KB
- ChangeColorMarkersInDmarkers.png 1,277 × 945; 923 KB
- Chiera-reference-center-gui.png 2,928 × 1,573; 1.77 MB
- Chimera-particles-center-gui.png 3,007 × 1,440; 987 KB
- Chimera-reference-center-gui-sta.png 2,884 × 1,493; 1.75 MB
- Chimera-volume-tracer-sta.png 722 × 1,034; 123 KB
- Chimera-volume-tracer.png 724 × 1,096; 147 KB
- Coarse-half-maps-alignment-configuration-sta.png 1,849 × 897; 184 KB
- Coarse-half-maps-alignment-configuration.png 3,016 × 1,360; 496 KB
- Computing environment gui standalone.png 1,394 × 1,614; 177 KB
- Concept of fiducial alignment.png 1,744 × 584; 197 KB
- Ctffind4 on unaligned TS.png 821 × 837; 621 KB
- CustomVideoMontageInline.png 897 × 958; 230 KB
- CustomVideoMontageSineRandom.png 900 × 961; 435 KB
- CustomVideoMontageSinuses.png 993 × 1,017; 271 KB
- DW DW and CTF.png 2,724 × 1,086; 3.7 MB
- Data organisation dynamo.jpg 267 × 402; 97 KB
- Dcm-gui-create-pre-binned-dmmpp.png 1,948 × 866; 229 KB
- Dcm-gui-create-pre-binned.png 1,954 × 875; 231 KB
- Dcm-gui-open-pre-binned-tomoslice-dmmpp.png 1,944 × 872; 239 KB
- Dcm-gui-open-pre-binned-tomoslice.png 1,954 × 881; 241 KB
- Dctutorial thumbnail gallery.png 1,918 × 1,204; 148 KB
- Default-mask-project-dcp-gui-sta.png 1,469 × 257; 26 KB
- Default-mask-project-dcp-gui.png 1,505 × 267; 27 KB
- Defocus left right.png 1,088 × 832; 111 KB
- Dhelp.png 1,866 × 1,570; 325 KB
- DipoleVesicleExampleCenter.png 353 × 242; 96 KB
- DipoleVesicleExampleDipoleActions.png 344 × 251; 103 KB
- DipoleVesicleExampleHideSpheresMenu.png 335 × 186; 55 KB
- DipoleVesicleExampleHideSpheresScene.png 344 × 251; 102 KB
- DipoleVesicleExampleNewDipoleMessage.png 237 × 192; 28 KB
- DipoleVesicleExampleNewDipolePopUp.png 404 × 279; 125 KB
- DipoleVesicleExampleNorth.png 353 × 251; 100 KB
- DipoleVesicleExampleScene.png 1,025 × 748; 458 KB
- DipoleVesicleExampleSecondDipole.png 344 × 251; 100 KB
- DipoleVesicleExampleSelectFirstDipole.png 344 × 251; 100 KB
- DipoleVesicleExampleSelection.png 521 × 279; 135 KB
- DipoleVesicleExampleWrongClick.png 372 × 242; 105 KB
- DpksliceBoxZoom.png 669 × 316; 119 KB
- Dpreview with region.png 1,500 × 876; 715 KB
- Dshow HIV1 raw.png 1,616 × 1,408; 3.01 MB
- Dtmshow.png 1,140 × 1,125; 533 KB
- DtmsliceBoxCreated.png 667 × 316; 77 KB
- DtmsliceBoxProjection.png 952 × 400; 259 KB
- DtmsliceBoxResetLimits.png 584 × 600; 207 KB
- DtplotBasic.png 1,120 × 840; 37 KB
- Dtview GUI 1.png 767 × 513; 39 KB
- Dtview GUI 3.png 666 × 448; 67 KB
- Dynamo console.png 764 × 423; 58 KB
- Dynamo graphical console.png 1,754 × 1,142; 125 KB
- Ec2CUDAAMI.png 1,500 × 893; 355 KB
- Ec2ChooseGPUInstance.png 1,350 × 743; 247 KB
- Ec2CreateKeys.png 1,500 × 893; 408 KB
- Ec2Dashboard.png 1,569 × 863; 416 KB
- Ec2FailedLaunch.png 1,500 × 1,066; 314 KB
- Ec2InstanceManager.png 1,350 × 743; 221 KB
- Ec2LaunchInstance.png 1,350 × 743; 327 KB
- Ec2LaunchInstanceInformation.png 1,500 × 1,045; 423 KB
- Ec2NonFreeTier.png 1,739 × 1,035; 467 KB
- Ec2NvidiaSelected.png 1,500 × 893; 353 KB
- Ec2Services.png 1,500 × 1,066; 528 KB
- Ec2SuccesfulLaunch.png 1,350 × 743; 262 KB
- Ec2Terminate.png 1,414 × 818; 346 KB
- Ec2runningInstancesDashboard.png 1,554 × 856; 406 KB
- Embo 2021 1.png 1,584 × 780; 69 KB
- Embo 2021 2.png 1,582 × 826; 117 KB
- Embo 2021 3.png 568 × 612; 52 KB
- Embo 2021 4.png 1,604 × 856; 164 KB
- Embo 2021 5.png 1,568 × 804; 320 KB
- Embo files.png 1,208 × 68; 40 KB
- Empty-sta-project-dcp-gui-sta.png 1,434 × 1,729; 182 KB
- Empty-sta-project-dcp-gui.png 1,454 × 1,746; 186 KB
- Environment-parameters-gui-sta.png 1,356 × 1,489; 133 KB
- Environment-parameters-gui.png 1,356 × 1,533; 138 KB
- ExampleTomoslice.png 1,918 × 1,123; 405 KB
- Example of healty FSC.png 745 × 540; 221 KB
- Exposition scheme HIV1.png 550 × 408; 13 KB
- FHVSymmetryAdhocMaskOverlay.png 533 × 401; 114 KB
- FHVSymmetryAdhocMaskPlot.png 498 × 374; 42 KB
- FHV zRandomized Parameters.png 783 × 736; 91 KB
- FHVsymmetryRecenteringOldMaskPlot.png 618 × 482; 59 KB
- FHVsymmetryRecenteringOldMaskSlice.png 631 × 473; 151 KB
- FSC mask Z.png 821 × 837; 26 KB
- FSC wrong phaseflip.png 945 × 685; 80 KB
- Fhv3dAntialiasing.png 1,022 × 597; 579 KB
- Fhv3dChimera.png 1,806 × 1,358; 507 KB
- Fhv3dDynamoTriangulation.png 1,250 × 1,125; 741 KB
- Fhv3dEmptyAxis.png 1,250 × 1,125; 132 KB
- Fhv3dFinal.png 1,056 × 690; 575 KB
- Fhv3dFinalTemplate.png 1,009 × 989; 185 KB
- Fhv3dHanddraw.png 1,050 × 933; 213 KB
- Fhv3dMapviewCentralSlice.png 1,481 × 1,125; 283 KB
- Fhv3dMapviewSym.png 1,409 × 1,011; 544 KB
- Fhv3dMapviewTemplate.png 1,480 × 1,125; 663 KB
- Fhv3dShifted.png 1,045 × 926; 649 KB
- Fhv3dSlice.png 1,065 × 933; 300 KB
- Fhv3dTrisurf.png 871 × 762; 207 KB
- Fhv3dTrisurfColormap.png 693 × 690; 267 KB
- Fhv3dTrisurfColormapCyan.png 706 × 683; 270 KB
- Fhv3dTrisurfLighting.png 720 × 678; 247 KB
- Fhv3dTrisurfNoEdges.png 908 × 697; 229 KB
- FhvBoxesModel.png 908 × 462; 210 KB
- FhvCloseUp.png 968 × 626; 406 KB
- FhvCloseUpClick.png 968 × 626; 406 KB
- FhvCloseUpClickCorrected.png 968 × 626; 406 KB
- FhvCreateSurfaceModel.png 1,180 × 1,125; 516 KB
- FhvDdbrowseFirstZ.png 1,526 × 1,233; 545 KB
- FhvDefinitionCrown.png 978 × 775; 422 KB
- FhvEnvironment.png 1,083 × 1,231; 148 KB
- FhvFirstAverage.png 1,153 × 1,125; 826 KB
- FhvFirstChimera.png 1,225 × 1,126; 450 KB
- FhvFirstParameters.png 1,192 × 1,125; 215 KB
- FhvHandDrawn.png 1,250 × 1,125; 247 KB
- FhvLocalizedChimera.png 1,224 × 1,125; 621 KB
- FhvLocalizedDview.png 1,510 × 771; 875 KB
- FhvLocalizedMapview.png 1,500 × 1,012; 466 KB
- FhvMeasureDistance.png 712 × 386; 210 KB
- FhvMembraneCompareWedges.png 1,480 × 1,125; 340 KB
- FhvMembraneMissingRandomized.png 1,153 × 1,125; 470 KB
- FhvMembraneMissingWedge.png 1,153 × 1,125; 447 KB
- FhvMembraneSketchDtplot.png 1,424 × 1,125; 189 KB
- FhvMembraneSlicesPostflipping.png 1,250 × 1,125; 519 KB
- FhvMembraneSlicesPreflipping.png 1,251 × 1,126; 520 KB
- FhvMembraneSurfaceEzplot.png 1,394 × 1,126; 235 KB
- FhvMontage.png 1,559 × 889; 626 KB
- FhvMontageActiveClick.png 1,064 × 1,125; 565 KB
- FhvMontageControls.png 1,455 × 792; 584 KB
- FhvOrientedSlicesX.png 1,250 × 1,125; 453 KB
- FhvOrientedSlicesZ.png 1,250 × 1,125; 520 KB
- FhvParametersLocalized.png 1,050 × 984; 185 KB
- FhvRandomizedFaintHintsMapview.png 1,395 × 916; 308 KB
- FhvSanityCheckZ.png 1,381 × 705; 646 KB
- FhvSelectedPoints.png 952 × 552; 296 KB
- FhvSketchFirst.png 1,251 × 1,125; 187 KB
- FhvSketchZ.png 1,250 × 1,125; 188 KB
- FhvSubboxAverage.png 1,444 × 722; 411 KB
- FhvSubboxOverlay.png 1,527 × 719; 368 KB
- FhvSubboxRefimentChimera.png 1,224 × 1,125; 324 KB
- FhvSubboxRefimentMapview.png 1,500 × 1,012; 414 KB
- FhvSubboxSelectDistance.png 1,500 × 1,012; 322 KB
- FhvSubboxSelectY.png 1,500 × 1,012; 313 KB
- FhvSubboxSelectZ.png 1,500 × 1,012; 424 KB
- FhvSubboxSketch.png 1,250 × 1,125; 322 KB
- FhvSurfaceControlPoints.png 1,466 × 1,150; 548 KB
- FhvSurfaceCreateTriangulation.png 1,278 × 1,126; 339 KB
- FhvSurfaceMarkCenter.png 916 × 759; 444 KB
- FhvSurfacePlotPoints.png 1,304 × 1,126; 344 KB
- FhvSurfaceSaveModel.png 1,501 × 906; 426 KB
- FhvSurfaceSubdivision.png 1,401 × 1,126; 370 KB
- FhvSurfaceSubdivisionMinimum.png 1,476 × 1,126; 386 KB
- FhvSurfaceWorkflowCreate.png 1,206 × 632; 245 KB
- FhvSymmetryDecentered.png 605 × 453; 173 KB
- FhvSymmetryGlobal.png 774 × 697; 79 KB
- FhvSymmetryGlobalWrong.png 505 × 379; 43 KB
- FhvSymmetryLocal.png 775 × 697; 77 KB
- FhvSymmetryLocalWrong.png 481 × 361; 42 KB
- FhvSymmetryRecenteredView.png 1,073 × 442; 588 KB
- FhvTemplateZ.png 1,501 × 773; 192 KB
- FhvTmsliceUpdates.png 1,500 × 906; 406 KB
- FhvTomoshowOnTheFly.png 1,412 × 1,075; 773 KB
- FhvTomosliceContrast.png 1,500 × 1,023; 442 KB
- FhvTomosliceOpen.png 1,500 × 904; 429 KB
- FhvZOrientedMapview.png 1,500 × 1,012; 290 KB
- FhvZOrientedParameters.png 1,192 × 1,126; 212 KB
- FhvZRandomizedChimera.png 1,224 × 1,126; 831 KB
- Fhv any computing enviroment.png 1,353 × 1,532; 159 KB
- Fhv computing enviroment.png 1,353 × 1,532; 153 KB
- Fhv environment.png 1,360 × 1,536; 191 KB
- FilamentAnchor1.png 1,500 × 907; 301 KB
- FilamentAnchor2.png 1,500 × 908; 285 KB
- FilamentControls.png 1,501 × 961; 299 KB
- FilamentSelectWorkflow.png 1,501 × 908; 358 KB
- FilamentSliceClickReady.png 1,250 × 1,125; 313 KB
- FilamentTypes.png 1,381 × 445; 182 KB
- FilamentUpdatePointsThroughSliceClick.png 1,499 × 885; 400 KB
- FilamentWorkflowStart.png 1,330 × 929; 170 KB
- Filament step 1.png 2,068 × 1,380; 447 KB
- Filament step 10.png 2,560 × 1,560; 391 KB
- Filament step 11.png 2,556 × 1,558; 576 KB
- Filament step 12.png 2,560 × 1,558; 725 KB
- Filament step 13.png 2,560 × 1,556; 406 KB
- Filament step 14.png 1,176 × 1,024; 316 KB
- Filament step 15.png 1,158 × 1,018; 299 KB
- Filament step 16.png 2,560 × 1,558; 603 KB
- Filament step 17.png 2,560 × 1,556; 784 KB
- Filament step 18.png 1,248 × 808; 183 KB
- Filament step 19.png 1,986 × 1,298; 469 KB
- Filament step 2.png 2,016 × 1,382; 425 KB
- Filament step 20.png 1,880 × 1,292; 440 KB
- Filament step 21.png 1,790 × 1,300; 427 KB
- Filament step 22.png 1,854 × 1,316; 298 KB
- Filament step 23.png 2,540 × 1,556; 760 KB
- Filament step 24.png 1,666 × 1,320; 164 KB
- Filament step 245.png 1,666 × 1,320; 164 KB
- Filament step 25.png 1,244 × 1,352; 120 KB
- Filament step 26.png 2,290 × 1,020; 509 KB
- Filament step 3.png 2,556 × 1,556; 469 KB
- Filament step 4.png 2,560 × 1,556; 640 KB
- Filament step 5.png 2,554 × 1,558; 564 KB
- Filament step 6.png 2,560 × 1,560; 602 KB
- Filament step 7.png 2,560 × 1,554; 412 KB
- Filament step 8.png 2,558 × 1,556; 338 KB
- Filament step 9.png 2,560 × 1,554; 762 KB
- Final-average-from-average-of-averages-sta.png 1,182 × 588; 453 KB
- Final-average-from-average-of-averages.png 1,925 × 971; 730 KB
- Final-result-from-refined-gold-standard-sta.png 2,933 × 1,466; 544 KB
- Final-results-from-refined-gold-standard-sta.png 4,276 × 2,433; 3.04 MB
- Final-sta-result-from-refined-gold-standard-sta.png 2,933 × 1,466; 1.34 MB
- Final FSC curve.png 892 × 617; 65 KB
- Final chimera HIV1.png 842 × 631; 487 KB
- First-order-distance-clusters-three-different-vlps.png 2,821 × 802; 84 KB
- First-per-tomogram-and-averages-of-averages-project-configuration-sta.png 1,915 × 887; 133 KB
- First-per-tomogram-and-averages-of-averages-project-configuration.png 2,988 × 1,272; 406 KB
- Folder structure HIV1.png 702 × 651; 123 KB
- Folder structure HIV1 v2.png 566 × 531; 90 KB
- Galactus TMor.png 847 × 408; 222 KB
- Galactus TMpre.png 867 × 419; 257 KB
- Galactus original.png 864 × 426; 386 KB
- Galactus pre.png 850 × 423; 389 KB
- General pipeline.png 1,866 × 450; 296 KB
- Geometrical-primitive-local-model-partition.jpeg 1,920 × 1,080; 340 KB
- Gmm-population-clasification-sta.png 4,365 × 639; 447 KB
- Gmm-population-clasification.png 4,390 × 660; 400 KB
- Gold beads quality.jpg 958 × 382; 324 KB
- H1 d h2.png 1,620 × 1,406; 1.35 MB
- HIV mask m.png 1,614 × 1,404; 846 KB
- HIV mask s.png 1,614 × 1,410; 801 KB
- Hello-world.png 661 × 715; 580 KB
- HivAverageCentered.png 1,466 × 729; 813 KB
- HivAverageLow.png 1,157 × 1,125; 685 KB
- HivBox.png 462 × 454; 197 KB
- HivClickDipole.png 1,431 × 475; 424 KB
- HivDDbrowseCentered.png 1,336 × 1,125; 690 KB
- HivDdbrowseInitial.png 1,371 × 1,125; 572 KB
- HivDisks.png 1,446 × 1,125; 575 KB
- HivDtmshow.png 1,246 × 1,125; 790 KB
- HivDtplotAll.png 1,250 × 1,125; 279 KB
- HivFinalSelection.png 1,500 × 903; 576 KB
- HivForgetDipoleOpening.png 793 × 670; 310 KB
- HivMapviewEvolution.png 1,480 × 1,125; 338 KB
- HivSelectModel.png 1,500 × 904; 508 KB
- HivSlicesEvolution.png 1,250 × 1,125; 224 KB
- HivWrongOnY.png 811 × 435; 167 KB
- HivWrongYReselect.png 835 × 384; 144 KB
- HoleClickerClickingGalleryDetail.png 186 × 165; 19 KB
- HoleClickerClickingHoverDetail.png 186 × 165; 20 KB
- HoleClickerClickingSwitchDetail.png 186 × 165; 19 KB
- HoleClickerClickingZoomOutDetail.png 186 × 165; 20 KB
- HoleClickerGUIWithPoints.png 560 × 543; 184 KB
- Image set section.png 835 × 774; 108 KB
- Imod alignment parameters.png 699 × 807; 261 KB
- Imod red box.png 875 × 813; 452 KB
- Imod residuals.jpg 969 × 1,040; 405 KB
- Initial-reference-project-configuration-sta.png 1,897 × 892; 132 KB
- Initial-reference-project-configuration.png 3,076 × 1,360; 497 KB
- Isosurface-final-structure-density-map-chimera-sta.png 4,248 × 1,672; 4.45 MB
- Isosurface-final-structure-density-map-chimera.png 4,248 × 1,672; 3.56 MB
- Error creating thumbnail: File with dimensions greater than 12.5 MPLast-average-coarse-half-maps.png 3,561 × 5,293; 4.6 MB
- Last-average-first-per-tomogram-average-sta.png 1,947 × 1,186; 2.07 MB
- Last-average-first-per-tomogram-average.png 1,966 × 1,210; 2.11 MB
- Last-average-for-coarse-half-maps-sta.png 5,027 × 824; 2.73 MB
- Last-average-for-coarse-half-maps.png 5,060 × 843; 2.04 MB
- Last-average-from-initial-reference-project-sta.png 1,920 × 957; 1.5 MB
- Last-average-from-initial-reference-project.png 1,920 × 957; 1.18 MB
- Last-average-second-per-tomogram-average-sta.png 2,027 × 1,217; 1.78 MB
- Last-average-second-per-tomogram-average.png 2,048 × 1,228; 1.8 MB
- LocalizedProjectionDtmshow.png 1,241 × 1,125; 792 KB
- LocalizedProjectionDtmshowBinnedZoomed.png 2,370 × 1,125; 1.01 MB
- LocalizedProjectionDtmshowFullZoomed.png 2,346 × 1,125; 1.53 MB
- LocalizedProjectionDtmshowRec.png 2,391 × 1,125; 1.48 MB
- LocalizedWBPBoxes.png 1,084 × 577; 269 KB
- LocalizedWBPMarkersAligned.png 1,454 × 1,125; 754 KB
- LocalizedWBPMarkersAlignedZoom.png 1,205 × 933; 231 KB
- LocalizedWBPSelectionBoxes.png 1,865 × 1,125; 1.11 MB
- LocalizedWBPSelectionBoxesZoom.png 1,866 × 1,125; 778 KB
- LocalizedWBPSelectionProjection.png 2,240 × 1,223; 1.39 MB
- LocalizedWBPSelectionProjectionConcurrent.png 1,980 × 1,213; 918 KB
- LocalizedWBPdmarkersRough.png 2,781 × 1,125; 978 KB
- LocalizedWBPdshow2d.png 2,261 × 1,125; 1.24 MB
- LocalizedWBPgoldbead1 3d.png 1,249 × 1,125; 710 KB
- LocalizedWBPgoldbead1 view.png 1,154 × 1,125; 467 KB
- LocalizedWBPlocalReconstruction1.png 1,153 × 1,125; 161 KB
- LocalizedWBPlocalReconstruction1Full.png 1,152 × 1,125; 820 KB
- LocalizedWBPlocalReconstruction1FullRecentered.png 1,152 × 1,125; 877 KB
- LocalizedWBPlocalReconstructionAllBinned.png 1,250 × 1,125; 179 KB
- LocalizedWBPlocalReconstructionAllFullCompare.png 2,254 × 1,070; 741 KB
- ManualAlignmentControlViewers.png 1,701 × 1,057; 255 KB
- ManualAlignmentControls.png 601 × 355; 49 KB
- ManualAlignmentDmarkers.png 1,455 × 1,125; 687 KB
- ManualAlignmentFitting.png 1,579 × 1,126; 767 KB
- ManualAlignmentReconstructorGUI.png 1,346 × 1,201; 187 KB
- ManualAlignmentSIRTWBP.png 1,550 × 716; 482 KB
- ManualAlignmentSetGoldBead.png 744 × 983; 511 KB
- Mapview handdraw slice zoom.png 1,250 × 1,125; 173 KB
- Mapview revolution mask chimera.png 1,315 × 1,126; 215 KB
- Mapview select slice for revolution.png 1,407 × 1,125; 309 KB
- Mask created by thresholding.png 1,250 × 1,125; 146 KB
- Mask handdraw montage.png 909 × 962; 211 KB
- MembraneDistanceFilters.png 560 × 420; 2 KB
- MembraneDistanceHistogram.png 560 × 420; 10 KB
- MembraneExampleCommandLineBothMeshes.png 1,167 × 875; 101 KB
- MembraneExampleCommandLineCroppingPositionsOnMesh.png 1,167 × 875; 60 KB
- MembraneExampleCommandLineCroppingPositionsOnMeshPerspective.png 1,167 × 875; 50 KB
- MembraneExampleCommandLinePointsControlAndTriangulation.png 1,167 × 875; 66 KB
- MembraneThicknessAverage.png 747 × 864; 41 KB
- MembraneWorkflowControlPoints.png 1,167 × 875; 44 KB
- MembraneWorkflowCropMesh.png 1,167 × 875; 46 KB
- MembraneWorkflowMesh.png 1,167 × 875; 52 KB
- MembraneWorkflowRefinedMesh.png 1,167 × 875; 80 KB
- MembraneWorkflowTableSketch.png 1,167 × 875; 43 KB
- MembraneWorkflowUserPoints.png 1,167 × 875; 41 KB
- Membrane step 1.png 2,072 × 1,386; 452 KB
- Membrane step 10.png 1,234 × 1,348; 275 KB
- Membrane step 11.png 2,398 × 1,358; 579 KB
- Membrane step 12.png 2,390 × 1,358; 611 KB
- Membrane step 13.png 2,388 × 1,370; 621 KB
- Membrane step 14.png 2,388 × 1,362; 472 KB
- Membrane step 16.png 2,378 × 1,354; 404 KB
- Membrane step 17.png 2,390 × 1,366; 615 KB
- Membrane step 18.png 2,380 × 1,356; 652 KB
- Membrane step 19.png 2,380 × 1,368; 682 KB
- Membrane step 2.png 1,984 × 1,356; 372 KB
- Membrane step 20.png 2,560 × 1,556; 834 KB
- Membrane step 21.png 1,662 × 1,330; 165 KB
- Membrane step 212.png 1,662 × 1,330; 165 KB
- Membrane step 22.png 1,352 × 1,286; 151 KB
- Membrane step 23.png 1,414 × 1,292; 896 KB
- Membrane step 3.png 2,558 × 1,558; 440 KB
- Membrane step 4.png 2,558 × 1,558; 875 KB
- Membrane step 5.png 2,560 × 1,560; 657 KB
- Membrane step 6.png 2,560 × 1,558; 664 KB
- Membrane step 7.png 2,560 × 1,560; 711 KB
- Membrane step 8.png 2,560 × 1,562; 698 KB
- Membrane step 9.png 2,560 × 1,556; 842 KB
- MercatorDview.png 1,172 × 1,125; 474 KB
- MercatorIsosurface.png 1,150 × 1,125; 388 KB
- MercatorRadiusClose.png 1,197 × 1,125; 573 KB
- MercatorRadiusFar.png 1,197 × 1,125; 442 KB
- MissingWedgeEstimationFactor0point1.png 1,572 × 846; 118 KB
- MissingWedgeVisualizationUnclear.png 1,572 × 846; 104 KB
- MissingWedgeminus45To60.png 1,572 × 846; 97 KB
- MissingWedgeminus60To45.png 1,524 × 852; 96 KB